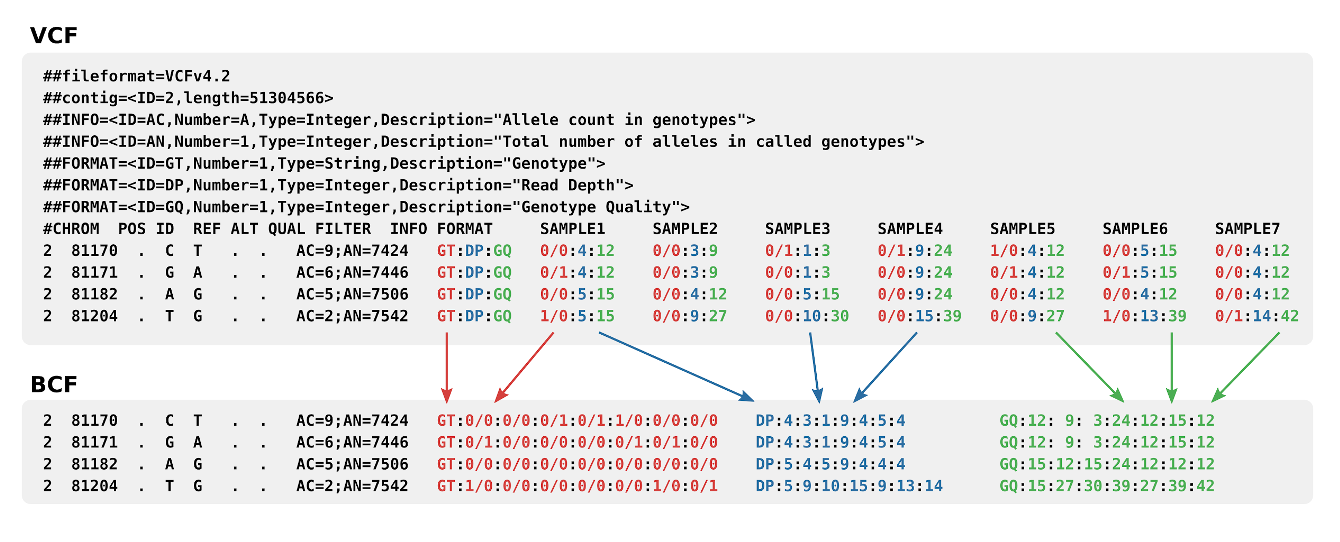
Comparison of selectivity and sensitivity of three SNP-calling tools... | Download Scientific Diagram
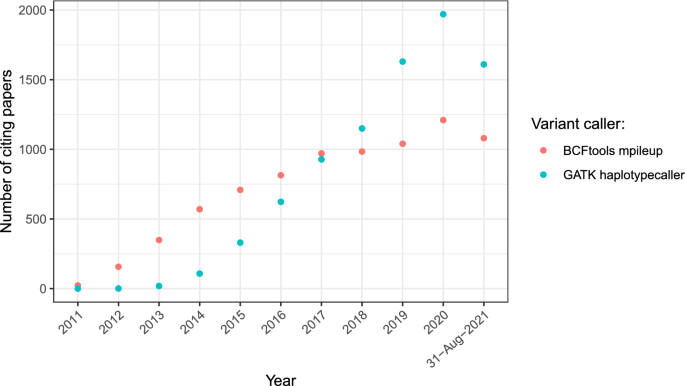
The evaluation of Bcftools mpileup and GATK HaplotypeCaller for variant calling in non-human species | Scientific Reports
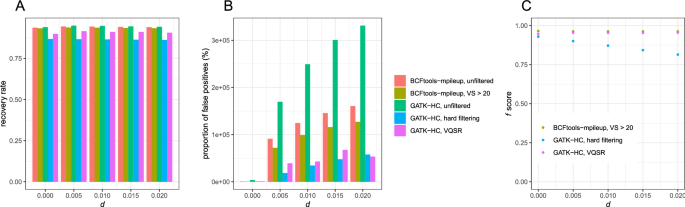
The evaluation of Bcftools mpileup and GATK HaplotypeCaller for variant calling in non-human species | Scientific Reports

Comparison of error rates of BCFtools/RoH and other existing methods as... | Download Scientific Diagram

bcftools view | bcftools tutorial on how to count the number of snps and indels in a vcf file - YouTube
GitHub - samtools/bcftools: This is the official development repository for BCFtools. See installation instructions and other documentation here http://samtools.github.io/bcftools/howtos/install.html
![PDF] BCFtools/RoH: a hidden Markov model approach for detecting autozygosity from next-generation sequencing data | Semantic Scholar PDF] BCFtools/RoH: a hidden Markov model approach for detecting autozygosity from next-generation sequencing data | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/9855a97fa2a760c4bfe74f5902f9de458f0a5a79/2-Figure1-1.png)